Unit 3 Chromosomes and Nucleic Acids
Key Unit Competence
To be able to describe the structure of a chromosome and how DNA is folded into achromosome
LEARNING OBJECTIVES
At the end of this unit, the learner will be able to:
• Describe the composition of chromosomes and the structure of nucleotides.
• Use of complementary base pairing to write the sequence for messenger RNA and the first
DNA codes for three base codons.
• Appreciate the importance of the presence of DNA in chromosomes.
• State how nucleotides pair.
• Describe the structure of DNA and RNA.
• Explain that the structure of the DNA molecule is described as a ladder twisted into a spiral.
• Draw the structure of DNA (6-10 base pair sequence).
• Explain the Watson-Crick hypothesis of the nature of DNA.
• Research on how Watson and Crick determined the nucleotide base pairing pattern.
• Outline the significance of telomeres in permitting continued replication.
• Acknowledge the role of telomeres in preventing the loss of genes and its relation to the
development of cancer.
• Distinguish between RNA and DNA.
• Describe the nature of genes.• Describe the structure of a genetic code.
INTRODUCTORY ACTIVITY
The genetic characters are passed from parents to offspring through the genes found in the
reproductive cells.
Conduct a research available resources to answer the following questions:
1. What is the relationship between a gene, DNA, and Chromosomes?
2. Draw and label the structure of a chromosome3. What do know about nucleic acids?
3.1 COMPOSITION OF CHROMOSOME
ACTIVITY 1
Aim: Observe the structure of chromosomes at prophase stage of mitotic cell division.
Materials Required:
1. Permanent slide of prophase stage of mitotic cell division.
2. Compound microscope
Background: Prophase is the first phase of mitosis. At this phase, no nucleolus is seen; the
chromosomes start condensing. Thread-like structure of chromosomes is seen under compoundmicroscope
Procedure:
1. Take the permanent slide.
2. Place it on the stage of the compound microscope.
3. Observe the prophase stage of mitotic cell division.4. Draw a well-labelled diagram of the structure of prophase stage chromatin.
Discussion:
1. Is there any difference between the structure of chromatin which you have observed and
what you have learned in the theory class?
2. What can you say about the nuclear membrane and spindle fibres in prophase stage observedunder a compound microscope?
Chromosomes are made of long aggregates of genes formed from condensed chromatin.
Chromatin is made up of DNA, proteins, RNA and other macromolecules. It is located in the
nucleus of a cell.
Deoxyribonucleic acid (DNA) is the storehouse of genetic information in the cell. A complete
set of an organism’s DNA is called a genome. And a gene is a segment of DNA that encodes
for a particular trait. Chromosomes are the structures that hold genes; they are made up of
strands of DNA tightly wrapped around histone proteins. Chromosome is basically composed
of three components—
(A) Nucleotides, (B) Histones proteins and (C) Non-histones proteins.
A. Nucleotides
The monomers that make up Deoxyribonucleic acid (DNA) and Ribonucleic acid (RNA) are
called nucleotides. Nucleotide has three components:
1. Pentose (five-carbon) sugar,
2. Nitrogenous (nitrogen-containing) base, and3. Phosphate group.
Pentose Sugar
Pentose sugar has five carbon atoms, which are numbered 1’ to 5’ respectively (Figure 3.1).
In DNA, the pentose sugar is deoxyribose: a hydrogen atom (H) is attached at the 2’ carbon
position. In RNA, the pentose sugar is ribose: hydroxyl group (OH) is attached at the 2’ carbonposition.
Figure 3.1: Structures of deoxyribose and ribose in DNA and RNA
Nitrogenous Bases
There are two classes of nitrogenous bases—Purines and Pyrimidines. Purines are ninemembered,
double-ringed structures (Figure 3.2). In these purines, the carbons and nitrogens
are numbered 1 to 9. There are two purines—Adenine (A) and Guanine (G). Pyrimidines
are six-membered, single-ringed structures (Figure 3.2). The carbons and nitrogens in these
pyrimidines are numbered 1 to 6. Pyrimidines are of three types—Thymine (T), Cytosine (C),and Uracil (U) (Figure 3.2).
Figure 3.2: Structures of nitrogenous bases in DNA and RNATable 3.1: Difference between DNA and RNA base composition
B. Histones Proteins
Histone proteins play an important role in organizing the physical structure of the chromosome.
They are most abundantly found in chromatin where they are wrapped around by DNA strands.
Moreover, they are small basic proteins with a net positive charge that assist their binding to
the negatively charged DNA (due to phosphate groups which are negatively charged).
In eukaryotes, there are five main types of histone proteins
(Figure 3.4). They are—H1, H2A, H2B, H3, and H4. H1
is loosely attached to the rest of the histone core proteins.
That is why H1 can be easily separated from the rest of the
histone proteins. And two each of H2A, H2B, H3, and H4
form core of eight histone proteins. These core proteins are
also called histone octamers. A strand of DNA measuring
147 bp segments wraps around this histone octamers for
about 1.7 times. Each nucleosome is connected by a strand
of DNA called linker DNA. For example, Human linkerDNA ranges from 38-53 bp long.
C. Non-histones Proteins
Excluding histone proteins, the rest of the proteins associated with DNA come under the
category of non-histone proteins. Non-histone proteins in so many ways are different from
histone proteins. Some of the differences are:
1. The number of non-histone proteins is much lesser than histone proteins.
2. Non-histone proteins are acidic proteins, which are negatively charged.
3. They play important role in the process of DNA replication, DNA repair, transcription
gene regulation, and recombination.
4. They vary in number and type from cell type to cell type within an organism at differenttimes in the same cell type, and from organism to organism.
3.1.1 The Importance of the Presence of DNA in Chromosomes
(a) Protection
The packaging of DNA in chromosomes helps in protecting DNA from being damaged.
(b) Conserve Space
If we take the DNA from all the cells in a human body and line it up, end to end, it would
form a strand 6000 million miles long! In order to store this very long important material,
DNA molecules are tightly packed around proteins called histones to make structures called
chromosomes.
The packaging of DNA in chromosomes helps in conserving space in cells. Approximately,about two metres of human DNA can fit into a cell that is only a few micrometres wide.
Figure 3.5: To better fit within the cell, long pieces of double-stranded DNA are tightlypacked into structures called chromosomes
(c) Control of Gene Expression
Chromatin is a complex of DNA and proteins that forms chromosomes within the nucleus
of eukaryotic cells. In its extended form, chromatin looks like beads on a string (Figure 3.5)
under the microscope. The beads are called nucleosomes, while the link between them is a
strand of DNA.
The packaging of DNA in chromatin form helps in controlling gene expression. Highly
compacted chromatin is not accessible to the enzymes involved in DNA transcription,
replication, or repair.
Chromatin has two main regions. The less condensed regions of chromatin are the regions
where active transcription takes place. This region is called euchromatin. On the other hand, the
condensed region of chromatin is where transcription is inactive or is being actively inhibitedor repressed. This region is called heterochromatin.
APPLICATION 3.1
Use the available resource to conduct research on the composition of chromosome. Thereafter,
answer the following questions:
1) What makes up a chromosome2) What is the difference between histone and non-histone proteins?
3.2 STRUCTURE OF NUCLEOTIDES
ACTIVITY 3.2
Use search engine to find out:
1. How the two strands in DNA join or stick together to form double stranded structure?2. Why adenine and guanine don not pair when the two strands join?
3.2.1 Nucleotide
A basic unit of nucleotide is made up of pentose sugar, a nitrogenous base, and a phosphate sugar.
However, a combination of only a pentose sugar and nitrogenous base, without phosphategroup, is called nucleoside.
In DNA and RNA, bases are covalently bonded to the 1’ carbon of the pentose sugar. The purine
and pyrimidines bases attached to pentose sugar from different positions of their nitrogen bases.
Purine bases use the 9th position of nitrogen to attach with 1’ carbon of pentose sugar, while
pyrimidine bases use the 1st position of nitrogen to attach with 1’ carbon of pentose sugar.
In both DNA and RNA, the phosphate group (PO4 2–) attaches to the 5’ carbon of pentosesugar.
Thus, by attaching phosphate group to a nucleoside yields a nucleoside phosphate or
nucleotide.
In DNA, the complex of deoxyribose, nitrogenous base and phosphate group is called
DNA nucleotide (a deoxyribonucleotide) (Figure 3.7), whereas in RNA, the complex of
ribose, nitrogenous base and phosphate group is called RNA nucleotide (a ribonucleotide)(Figure 3.6).
3.2.2 Phosphodiester Bond
Two nucleotides are covalently joined together by a bond called phosphodiester bond
(Figure 3.8). In phosphodiester bond, the phosphate group, which is attached on 5’ of
one nucleotide, forms a bond with the 3’ carbon of another nucleotide. In this way, many
phosphodiester bonds are formed in between sugar and phosphate groups. The repeated sugarphosphate-
sugar-phosphate backbone is a strong one. Because of this strong backbone, DNAand RNA are stable structures.
5′ end
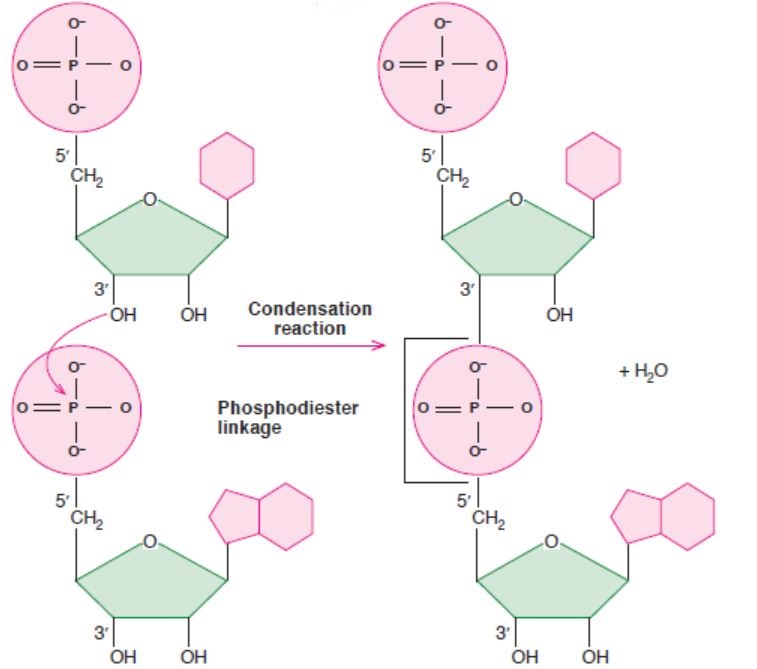
3′ end
Figure 3.8: A figure showing phosphodiester bond formation (between two ribonucleosides)
3.2.3 Polarity
Polynucleotide chains have polarity. On one end, there is a 5’ carbon with a phosphate group
(PO4
2–). On the other end, there is a 3’ carbon with a hydroxyl group (OH) on it (Figure 3.8).
The ends of polynucleotide are frequently referred to as the 5’ end and the 3’ end.
3.2.4 Chargaff ’s Rules: The Rules of Base Pairing
Erwin Chargaff’s rules state that DNA of all organisms should have a 1:1 ratio of pyrimidine and
purine bases. Thus, the amount of adenine (A) is equal to that of thymine (T); and the amount
of guanine (G) is equal to that of cytosine (C). This equivalence of purine and pyrimidine bases
is known as Chargaff ’s rules. This pattern is found in both strands of DNA.Table 3.2: Difference between purine and pyrimidine
The specific base pairing of A-T bases and G-C bases is called complementary base pairs. For
example, if one strand of DNA sequence is 5’-ATATCCGGAT-3’, then the opposite strand of
DNA sequence will be 3’-TATAGGCCTA-5’. Thus, by using the rules of base pairing, once
we have the sequence of at least DNA strand, we can find out the opposite base sequence ofthat DNA.
APPLICATION 3.2
1) Complete the sentence with the correct word:
(a) ............................ is a segment of DNA that encodes for traits.
(b) The two Purines are ............................... and ................................... .
(c) Uracil is present in .................................. .
(d) Two nucleotides are covalently joined by ..................................... .
(e) A-T bases and G-C bases are called ................................... base pairs.
2) What is the maximum number of hydrogen bonds in a length of DNA containing 700 basepairs?
3.3 STRUCTURE OF NUCLEIC ACIDS—DNA AND RNA
ACTIVITY 3.3
Aim: Design the structure of DNA molecule and complimentary
base pairing using plastic
model shapes or homemade kits.
Materials Required:
1. Plastic models of pentose sugars, phosphate groups, nitrogenous bases (A, G, T, and C).
Or
2. Homemade kits of pentose sugars, phosphate groups, nitrogenous bases (A, G, T, and C).
Procedure:
1. Before you start the activity, study the composition of chromosome (DNA) properly.
2. By using the plastic models, construct the structure of DNA.
3. Or in the same way, by using the plastic models, make the complementary base pairs of
nucleotides (A, G, T, and C).
4. Once the construction of the structure of DNA and the complementary base-pairing are
ready, give a presentation to the class.
Note: 1. Remember the Chargaff ’s rule of base pairing concept.2. Remember the structure of DNA.
In the structure of DNA, the strong electronegative atom is the Oxygen (O) and Nitrogen (N),
while H atom has positive charge. In the structure of DNA (Figure 3.10), thymine and adenine
have two hydrogen bonds; while guanine and cytosine have three hydrogen bonds. Hydrogen
bonds or interactions play a very important role in binding the bases of the opposite strands in
the DNA. Though RNA is not the genetic material in most of the cases, both single-stranded
and double-stranded RNAs are the genomes of certain viruses. RNA double-stranded molecules
show structural similarity to that of double-stranded DNA molecules. The similarities are:
(a) Both have anti-parallel strands.
(b) Both have sugar-phosphate backbones on the outside of helical molecule.
(c) In both the cases, in the middle of the helix, a complementary base pairing is formed byhydrogen bonds.
Figure 3.10: Structure of DNA [See the hydrogen bonds between the four bases (T-A, G-C)]
Figure 3.11: Chemical structure of RNA
APPLICATION 3.3
1) Draw the structure of a DNA sequence made of 3 pairs of nucleotides
2) Compare DNA and RNA
3.4 THE WATSON-CRICK HYPOTHESIS OF THE NATURE OF DNA/THESTRUCTURE OF DNA
ACTIVITY 3.4
In 1953, James D. Watson, an American molecular biologist, and Francis H.C.Crick, a British
molecular biologist, proposed a model for the physical and chemical structure of the DNAmolecule.
Make research from available resource to explain why DNA is described as a Twisted LadderStructure.
In 1953, James D. Watson, an American molecular biologist, and Francis H.C.Crick, a British
molecular biologist, proposed a model for the physical and chemical structure of the DNA
molecule. Today, their model is known as double helix model of DNA or simply the Structureof DNA.
The main features of Watson and Crick double helix model (Figure 3.12) of DNA are:
1. Two polynucleotide chains wind around each other in a right-hand double helix(Figure 3.12).
2. The two polynucleotide chains run side-by-side in an antiparallel fashion. This means
that one strand of DNA will orient itself in a 5’ -3’ direction, whereas, the other strand
will orient itself alongside the first one in a 3’-5’ direction. In this way, the two strandsare oriented in opposite directions (Figure 3.13).
3. On one hand, the sugar-phosphate backbones lie outside of the double helix. On the
other hand, the bases orient themselves toward the central axis of the double helix
structure.
The bases of one strand are bonded with the bases of the other strand of double helix
by hydrogen bonds. These bonds are weak chemical bonds. Since hydrogen bonds are
relatively weak bonds, the two strands can be easily separated by heating the DNA. The
bonding of these bases in the double helical structure follows the Chargaff ’s base pairingrules. For example—Adenine (A) will form two hydrogen bonds with Thymine (T).
Similarly, Guanine (G) will form three hydrogen bonds with Cytosine (C). This specificbase paring is called complementary base pairing (Figure 3.12).
4. The distance between adjacent bases is 0.34 nm in the DNA helix. A complete turn of
the helix takes 3.4 nm. One complete turn, which is 360° turn, accommodates 10 base
pairs (bp). And the diameter of the helix is 2 nm (Figure 3.13).
5. There are major and minor grooves in the double helix. The two sugar-phosphate
backbones of the double helix are not equally spaced from one another along the helical
axis, because of the way the bases bind with each other. As a result, there is an unequal
size of grooves between the backbones. The wider groove is called major groove; rich in
chemical information. The narrower groove is called minor groove; less rich in chemical
information (Figure 3.13).
3.4.1 DNA is also Described as a Twisted Ladder Structure
A typical ladder has two long wooden or metal side strands or pieces between which a series of
rungs or bars are set in suitable distances (Figure 3.14). In the structure of DNA, the pentose
sugars and phosphate groups make up the “long two side strands or pieces” of a typical ladder.
And the A-T and G-C base pairs which are bonded by hydrogen bonds make up the “rungs
or bars”of a typical ladder (Figure 3.15).
But unlike a typical ladder which is straight, the two strands of DNA are twisted into spiral.
Scientists call this a double helix. DNA also folds and coils itself into more complex shapes. The
coiled shape makes it very small. In fact, it is small enough to easily fit inside any of our cells. If
a DNA from a cell is unfolded, it would stretch out to a length of about six feet. The structural
twisted nature of DNA has been attributed to enhance its stability and strength. Thus, for thesesimple similarities with a typical ladder, DNA is also referred to as a twisted ladder structure.
APPLICATION 3.4
1. Complete the sentence with the correct word
(a) Watson and Crick proposed the model of .................................... .
(b) Enzyme ..................................... maintains the length of telomere.
(c) ...................................... can be used to cure cancer.
(d) ..................................... bonds are seen in both DNA and RNA.
(e) ...................................... directs synthesis of proteins in the body.
2. Differentiate the structure of DNA to that of real ladder.
3. What would be the side effect of untwisting the DNA
3.5 SIGNIFICANCE OF TELOMERE IN PERMITTING CONTINUEDREPLICATION
3.5.1 What is Telomere?
A telomere is a region of repetitive nucleotide sequences at each of a chromosome. It protects
the end of the chromosome from being deleted or from fusion with neighbouring chromosomes.
In vertebrates, the repetitive sequence of nucleotides in telomeres is TTAGG. In humans, thissequence is repeated about 2500 times.
3.5.2 How is the Length of Telomere Maintained?
In the process of DNA replication, the length of chromosome (precisely the length of telomere) is
shortened by about 25-200 bases per replication cycle. Elizabeth Blackburn and Carol W. Greider
have shown that the enzyme telomerase maintains the length of chromosome by adding telomere
repeats (TTAGGG) at 3’ end overhang, which serves as template to previous DNA replication
(Figure 3.17). The complementary sequence (5’ end) to this newly synthesized telomere is thenadded by the regular replication machinery.
3.5.3 Significance of Telomere in Replication
1. Telomeres help in organising the chromosomes in the nucleus of the cell.
2. Telomeres protect loss of important genes: During DNA replication, the chromosomes are shortened
by about 25-200 bases per replication. If this process of shortening the chromosomes continues,
there will be loss of important genes. However, fortunately, the ends of the chromosomes are
protected by telomeres; and telomeres are non-coding regions. Thus, even if there is loss at the
tip of chromosomes in every round of replication, the loss of telomeres (non-coding regions)doesn’t affect the important genes
3. Telomeres prevent the end of chromosomes from fusing with its neighbouring chromosomes.
Telomerase has an associated RNA that complementsthe 3’ overhang at the end of the chromosome.
The RNA template is used to synthesize the complementary strand.
Telomerase shifts and the process is repeated.
Primase and DNA polymerase synthesize the complementary strand.
Figure 3.17: Synthesis of telomeric DNA by telomerase
Telomeres and Ageing
Telomeres are thought to be related to ageing. Newborn babies are reported to have telomeres
ranging from around 8000 to 13,000 base pairs. These base pairs tend to decline by around
20-40 every year. Thus, by the time someone is 40-year-old, they could have lost up to 1600 base
pairs from their chromosomes. However, no significant shortening of telomeres is observed in
old people. It has been observed that telomerase is typically active in germ cells and adult stem
cells, but is not active in adult somatic cells.
Telomeres and Cancer
Cancer cells are characterized by their rapid and uncontrollable division of cells. These
cells have active telomerase to help them divide uncontrollably and become immortal. In
the absence of telomerase, the cancer cells would become inactive and would stop dividing
resulting into death of the cancer cells. Cancer therapies can take advantage of this concept by
designing drugs that can inhibit telomerase activity, thereby killing the cancer cells. Telomere
biology is an important aspect of human cancer. Many scientists are hoping and working
hard to understand the best way to use anti-telomerase therapy and advance the treatmentof cancer.
3.5.4 Major Differences between DNA and RNA
The three major structural differences of RNA from that of DNA are:
1. RNA contains ribose sugar instead of 2’-deoxyribose. It means that ribose has a hydroxyl
group (OH) at the 2’ position, whereas, deoxyribose has hydrogen (H) at 2’ position in
pentose sugar.
2. RNA has Uracil (U), whereas DNA has thymine (T).
3. Unlike DNA, which consists of two polynucleotide chains, in most cases, RNA is foundin a single polynucleotide chain.
Table 3.3: Differences between DNA and RNA
APPLICATION 3.5
Complete the sentence with the correct word
(i) Watson and Crick proposed the model of .................................... .
(ii) Enzyme ..................................... maintains the length of telomere.
(iii) ...................................... can be used to cure cancer.
(iv) ..................................... bonds are seen in both DNA and RNA.(v) ...................................... directs synthesis of proteins in the body.
3.6 NATURE OF GENES
ACTIVITY 3.6
Just like a person has his/her own character and personality, genes also have their unique
characters or nature. By nature we mean the inherent character or basic constitution of a gene.
So, find out the nature of genes from the internet. Make a report on the same and present tothe class.
1. A complete set of an organism’s DNA is called genome. A gene is a segment of DNA
that encodes for a particular trait. For example—black hair, brown hair etc.
2. Chromosomes are the structures that hold genes; they are made up of strands of DNA
tightly wrapped around histone proteins.
3. Genes are located on the chromosomes.
4. In the chromosome, a gene is found in a pair or
alternative forms called alleles. An allele is one of
two or more versions of the same gene or gene locus.
Two alleles for each gene, one from each parent, are
passed on to offspring. Homozygous pair refers to two
of the same alleles (Figure 3.18); and heterozygous
pair refers to two different alleles.
5. Each gene allele occupies a specific position in each
chromosome called locus (plural- loci).
6. Alleles are either dominant or recessive. Dominant
allele will be expressed wherever it is present, even if
it is paired with recessive allele. But recessive allele is
expressed only when it is paired with another recessive
allele.
7. When two or more alleles are present in a gene, the
condition is called multiple alleles. Example: Human blood types. The ABO Blood Type
in human beings is determined by three alleles. IA, IB, i. Both IA and IB are codominant
alleles. They are dominant to the allele, “i.” Allele “i” is recessive (Table 3.4).Table 3.4: Human blood type
8. The gene may change its phenotypical (trait) expression due to sudden change in its
genetic composition. The changed gene is known as mutant gene. The phenomenon of
change in genetic composition is known as mutation.Figure 3.19: Mutation in a DNA sequence. [T base is replaced by C base]
9. Genes duplicate themselves very accurately by DNA replication. DNA replication is the
process of producing two identical DNA replicas from one original DNA molecule duringcell cycle. It occurs in all living organisms and is the basis for biological inheritance.
10. Central Dogma:
The central dogma of molecular biology is an explanation of the flow of geneticinformation, from DNA to RNA, to make a functional protein within a biological system.
Figure 3.20: Central Dogma-information flow in a biological system
11. Split Genes:
In most eukaryotes, the genes are not continuous. Rather, the exons, the coding regions, in
the m-RNA are interrupted by several distinct units of non-coding regions called intron.
Since, the exons are split by introns, such genes are called split genes or mosaic genes or
discontinuous genes. The intron regions are removed in the latter stage of transcriptionby a process called splicing. However, introns are absent in prokaryotes.
Figure 3.21: A diagram showing m-RNA regions of exons and introns
12. Genetic Code:
In the process of translation, ribosome reads the sequence of the m-RNA in a group of
three nucleotides called codons or nucleotide triplets. Thus, the genetic code is the set
of rules by which information encoded in the form of codons or nucleotide triplets in
the m-RNA is translated into proteins by living cells using ribosome machinery. Each
codon specifies a particular amino acid with some exceptions.
13. One Gene/One-Polypeptide Hypothesis:
George Beadle and Edward Tatum came up with the idea that each gene encodes the
structure of one enzyme. This idea was called the one-gene/one-enzyme hypothesis.
However, presently it is known that many enzymes have multiple polypeptide subunits,
and each subunit is encoded by a separate gene. This relationship is now referred to as
the one gene/one-polypeptide hypothesis.
APPLICATION 3.6
1. What is meant by:
a) Codon
b) Central Dogma
2. How would you summarize the hypothesis of George Beadle and Edward commonlyknown as one gene/one-polypeptide hypothesis?
3.7 GENETIC CODE
ACTIVITY 7
1. Discuss the following in the class.
The genetic code (codons) in m-RNA code for amino acids; and there are 64 possible codons
(sense and nonsense codons). But there are only 20 standard amino acids. How is it possible
that there are excess codons present to code for only 20 amino acids?2. Make a report on it and present it to the class.
Genetic Code
The genetic code is the set of rules by which information encoded in genetic material (DNA
or RNA sequences) is translated into proteins (amino acid sequences) by living cells using
ribosome machinery. In other words, the genetic code is a set of rules that specify how the
nucleotides sequence (ATGC) of an m-RNA is translated into the amino acid sequence of a
polypeptide chain.
The Structure of Genetic Code
The structure of genetic code is related to a series of exciting discoveries.
It was George Gamow (1954), a physicist, who argued that since there are only 4 bases and
if they have to code for 20 amino acids, the code should constitute a combination of bases.
In order to code for all the 20 amino acids, he suggested that the code should be made up
of three nucleotides (triplet code). The permutation and combination of three nucleotides
43 (4 × 4 × 4) would generate 64 codons. Proving that codon was triplet (i.e., three nucleotides)
was quite a challenging task. But the chemical method developed by Har Govind Khurana
for synthesizing RNA molecules with defined combinations of bases (homopolymers and
copolymers), and Marshall Nirenberg’s cell free system for protein synthesis finally helped
the genetic code to be deciphered. In the 1968, both of them, Marshall Nirenberg and Hare
Gobind Khurana along with Robert Hollye were awarded Nobel Prize in Physiology and
Medicine. Finally, a checker board for genetic code (Figure 3.22) was prepared which is asfollows.
Figure 3.22: Genetic Code: The first, second and third bases as read from 5’ to 3’ direction
constitute the triplet code in RNA. The codon AUG specifies methionine and is usually the
starting point for protein synthesis. The word ‘stop’ indicates codons serving as signals to
terminate protein synthesis. For each amino acid more than one codon have been identified.
It would be clear from the Figure that while the first and second bases remain the same for a particularamino acid, the third base can be different
APPLICATION 3.7
1. Complete with the correct word:
(a) The phenomenon of change in genetic constitution is called ......................
(b) Two of the same alleles result in ............... pair.
(c) Alleles can be ......................... or ............................ .
(d) The sense codon AUG, is a ............................ codon.2. What is meant by non-sense codons? State them with reference to eukaryotes.
3.8 SUMMARY
• Chromosome is composed of three main components: Nucleotides, histones proteins,
and non-histones proteins.
• Nucleotide is subdivided into pentose sugar, nitrogenous bases, and phosphate groups.
• The presence of DNA in chromosomes is important for three main reasons—Protection
from damage, conserve space, and control of gene expression.
• Nucleotide is made up of pentose sugar, nitrogenous bases, and phosphate groups;
whereas, nucleoside is made up of pentose sugar and nitrogenous bases.
• Phosphodiester bond connects the phosphate group, which is attached on 5’ of one
nucleotide, with the 3’ carbon of another nucleotide. This bond is a strong bond. That
is why DNA is a stable structure.
• Polynucleotide chains have polarity. On one end, there is a 5’ carbon with a phosphate
group. On the other end, there is a 3’ carbon with a hydroxyl group on it.
• Chargaff ’s rules state that DNA of all organisms should have a 1:1 ratio of purine
(A, G) and pyrimidine (T, C) bases. The specific base pairing of A-T bases and G-C bases
is called complementary base pairs.
• In 1953, Watson and Crick proposed the double helix structure of DNA.
• The two strands of DNA are anti-parallel; the bases on both strands are bonded by
hydrogen bonds in line with Chargaff ’s rules. DNA has major and minor grooves.
• DNA is also described as twisted ladder structure.
• RNA has a hydroxyl group at 2’ carbon of pentose sugar. It has a uracil base instead of thymine.
• Unlike DNA, RNA is not the genetic material of many organisms except for few viruses.
• DNA is double stranded while RNA is normally single stranded; DNA transfer genetic
material while RNA is involved directing the synthesis of proteins.
• A telomere is a region of repetitive nucleotide sequences at each of the tip of chromosomes.
• Telomere protects important genes from being deleted, and thus allows a continued
replication.
• Telomere regions are synthesized by a telomerase enzyme.
• Telomeric regions are important in ageing and cancer treatment.
• A gene codes for a specific trait.
• A particular gene can be present in two versions called alleles. When more than two
versions of gene are present, it is called multiple alleles.
• Alleles can either be dominant or recessive.
• Genes duplicate themselves through the process of DNA replication.
• Genes are copied from DNA to RNA through a process called transcription.
• Message in the m-RNA is translated into proteins through a process called translation.
• Many enzymes have multiple polypeptide subunits, and each subunit is encoded by a
separate gene. This relationship is called one gene/one-polypeptide hypothesis.
• It is the set of rules by which information encoded in genetic material (DNA or RNA
sequences) is translated into proteins (amino acid sequences) by living cells.
• Out of 64 codons, 61 codons are sense codons and 3 codons are non-sense codons.
• A codon is made up of three nucleotides or triplets.• Genetic code is almost universal; it shows degeneracy.
3.9 GLOSSARY
• Alleles: An allele is one of two or more versions of the same gene or gene locus.
• Central dogma: The flow of genetic information, from DNA to RNA, to make a functional
protein within a biological system.
• Chargaff ’s rule: It is a rule that states that DNA of all organisms should have a 1:1
ratio of pyrimidine and purine bases. Thus, the amount of adenine (A) is equal to that
of thymine (T); and the amount of guanine (G) is equal to that of cytosine (C).
• Chromosomes: They are the structures that hold genes; they are made up of strands of
DNA tightly wrapped around histone proteins.
• Gene: A gene is a segment of DNA that encodes for a particular trait.
• Genetic code: It is the set of rules by which information encoded in the form of codons or
nucleotide triplets in the m-RNA is translated into proteins by living cells using ribosome
machinery.
• Histone proteins: They are proteins that play an important role in organizing the physical
structure of the chromosome.
• Multiple alleles: The condition where two or more alleles are present in a gene. The
ABO Blood Type in human.
• Nucleoside: A combination of only a pentose sugar and nitrogenous base, without
phosphate group, is called nucleoside.
• Nucleotides: The monomers that make up deoxyribonucleic acid (DNA) and ribonucleic
acid (RNA) are called nucleotides. Nucleotide has three components:—
Pentose (fivecarbon)
sugar, Nitrogenous (nitrogen-containing) base, and Phosphate group.
• Phosphodiester bond: It is a chemical bond where the phosphate group, which is attached
on 5’ of one nucleotide, forms a bond with the 3’ carbon of another nucleotide. Many
phosphodiester bonds are formed in between sugar and phosphate groups.
• Split genes: The exons, the coding regions, in the m-RNA are interrupted by several
distinct units of non-coding regions called intron. Since the exons are split by introns,
such genes are called split genes or mosaic genes or discontinuous genes.
• Telomerase enzyme: It is an enzyme which maintains the length of chromosome by
adding telomere repeats (TTAGGG) at 3’ end overhang, which serves as template on
previous DNA replication.
• Telomere: A telomere is a region of repetitive nucleotide sequences at each of a
chromosome.
END UNIT ASSESSMENT 3
I. Choose whether the following statements are True (T) or False (F)
1. In DNA, the pentose sugar is ribose.
2. RNA has a hydrogen atom at 2’ carbon position.
3. Pyrimidine is a single-ringed structure.
4. DNA contains adenine, thymine, guanine, and cytosine.
5. Out of these five histone proteins, H1 is loosely attached to the rest of the histone
core proteins.
6. Non-histone proteins are acidic proteins.
7. Purine bases use its 9 position nitrogen to attach with 1’ carbon in pentose sugar.
8. DNA is left-handed double helix.
9. UAG, UGA and UAA nucleotides are stop codons.
10. In genetic code, degeneracy means degeneration of DNA.
II. Multiple Choice Questions
1. Codon is a group of
(a) 2 nucleotides (b) 3 nucleotides
(c) 4 nucleotides (d) 5 nucleotides
2. Newborn babies have telomeres ranging from around
(a) 8,000 to 13,000 base pairs (b) 8,000 to 16,000 base pairs(c) 8,000 to 12,000 base pairs (d) None of these
3. Split genes are
(a) Genes with splitting chromosomes
(b) Genes separated from one another
(c) Genes where exons are interrupted with introns
(d) Genes where introns are interrupted with exons
4. Mutant gene is
(a) A gene with different mother genes
(b) A gene where nucleotide sequence has changed due to mutation
(c) A gene where different genes exist together
(d) A gene of different shape and size
5. Splicing is process of
(a) Removing exons (b) Removing introns
(c) Removing coding genes (d) Removing DNA
6. Recessive allele will express only when
(a) It occurs with dominant alleles (b) It occurs with other recessive allele
(c) It is absent (d) It is present with proteins
7. DNA Replication is the process of
(a) Copying DNA from RNA (b) Copying DNA from proteins
(c) Copying DNA from DNA (d) Copying DNA from ribosome
8. Nitrogenous bases of the two strands of DNA are linked with
(a) Hydrogen bonds (b) Covalent bonds
(c) Ionic bonds (d) Phosphodiester bonds
9. DNA does not have
(a) Adenine (b) Cytosine
(c) Guanine (d) Uracil
10. Gene codes for
(a) Polypeptides (b) Blood
(c) Specific trait (d) Specific genome
III. Long Answer Type Questions
1. In your own words, describe the composition of chromosomes.
2. List at least three differences between the structures of DNA and RNA.
3. Why is DNA important in chromosomes?
4. What is telomere? Give the significance of telomere in replication and its importance
in cancer treatment.
5. Describe structure of a Genetic code.
6. In your own words, explain why the structure of DNA is described as a ladder twisted
into a spiral.
7. Draw the structure of DNA having at least 6 base pair sequence.
8. How did Watson and Crick determine the nucleotide base pairing pattern? Explain
in your own words.
9. In your own words, describe the nature of genes.
10. (i) Identify the structure shown in figure.
(ii) Write the measurement (distance) of the parts marked (1), (2), (3), (4) and (5).
(iii) How many H-bonds are there at the place marked (6)?
(iv) How many different forms of the shown structure have been reported to occur
in the living organisms? Give their names.
(v) Which of them has/have left handed spiral and which of them right handed
spiral?(vi) Mention any 2 other special features of the form having left handed spiral.
11. Explain the role of nucleic acids in detecting HIV-AIDS. Describe NAT and also tell
why NAT is not suitable for detecting ultra-law HIV-I DNA and RNA within hostcellular compartments.
